Main Second Level Navigation
Crowdsourcing Cancer Challenge
Jovana Drinjakovic
An international consortium — including University of Toronto professors Quaid Morris and Paul Boutros – has launched a global crowdsourcing competition to find new tools for researching cancer.
Research groups from Canada, the U.S. and the U.K. have joined forces to create a public contest in software development in order to address one of the greatest challenges in cancer biology. The contestants will analyze vast amounts of DNA sequence data to identify genetically distinct groups of cells within tumours that are often the reason why therapy fails.
Called the ICGC-TCGA DREAM Somatic Mutation Calling Heterogeneity (SMC-Het) Challenge, the competition is now open and runs until May 2016.
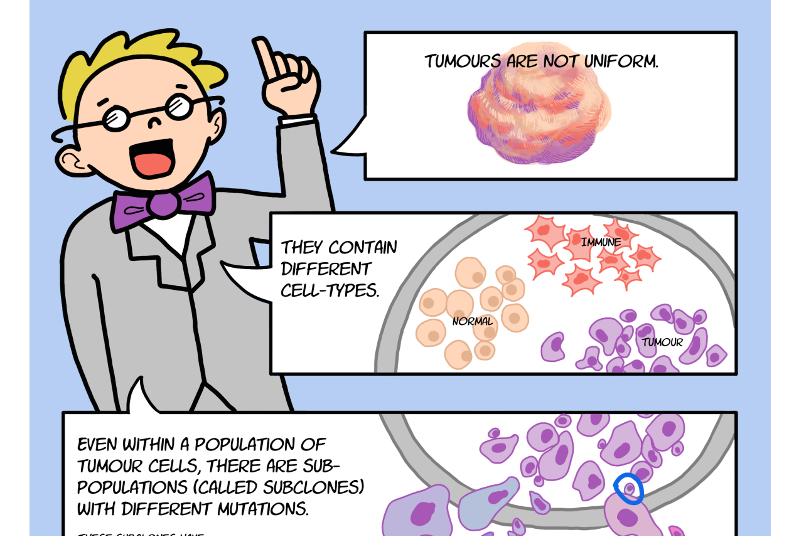
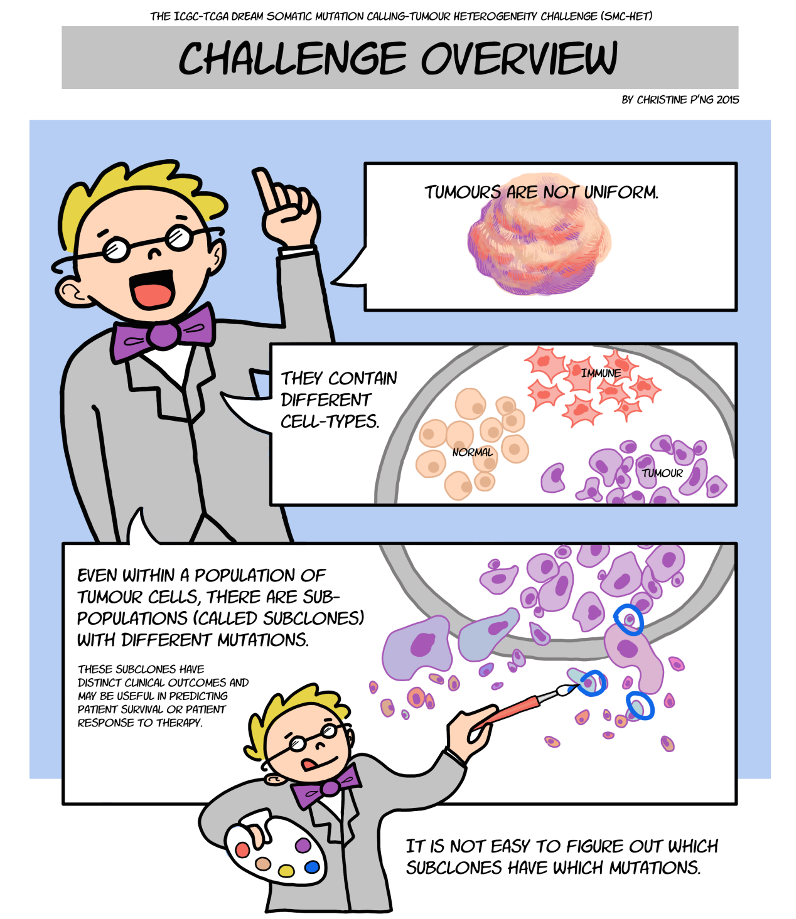 Cancer remains a leading cause of death throughout the world because of its ability to evade even the best available treatments. Recent advances in DNA sequencing reveal that tumour cells don’t all share the same DNA – rather some have acquired unique mutations that cause them to respond differently to therapy and spread around the body at different rates. To find an effective treatment, we first need an understanding of the many different types of cancer cells present in each patient.
Cancer remains a leading cause of death throughout the world because of its ability to evade even the best available treatments. Recent advances in DNA sequencing reveal that tumour cells don’t all share the same DNA – rather some have acquired unique mutations that cause them to respond differently to therapy and spread around the body at different rates. To find an effective treatment, we first need an understanding of the many different types of cancer cells present in each patient.
“We know that cancers are made up of many different populations of cells, known as ‘subclones’, and understanding the relationships between these subclones is critical in developing successful long-term treatments,” said Dr. David Wedge, Staff Scientist at the Wellcome Trust Sanger Institute in Cambridge, U.K. and one of the co-organizers.
The new ease of collecting patients’ genomic information also poses a challenge of how best to analyze the growing mountain of data.
“There’s been an explosion of computational methods for teasing cancer subclones apart, each using different strategies and making different assumptions. We wanted to organize this competition as a way of providing independent assessment of which approaches work the best,” said Morris, who is also a professor in U of T’s Donnelly Centre and the Department of Molecular Genetics.
The competition merges the efforts of the world’s largest cancer genome sequencing consortia with DREAM Challenges and Sage Bionetworks - the world-leading organizations in community-based open science that championed crowdsourcing as way of finding answers to some of the biggest questions in biomedicine.
“Crowdsourcing has provided a way for researchers across the world to collaborate on solving problems together. As a whole, the SMC Challenges have provided focus on specific problems that are understudied, and provided an easy way for scientists from other fields, and citizen scientists, to get involved in solving key problems in cancer genomics,” said Boutros, a scientist at the Ontario Institute for Cancer Research and a professor in the Department of Pharmacology and Toxicology and Department of Medical Biophysics.
The organizers have created a set of 50 realistic tumours with distinctive life histories and evolutions. Contestants will create tools for detecting cancer subclones in the cloud, donated by Google and using the Google Compute Engine that will be run in Galaxy, a widely used open-source platform in biomedical research. This will allow researchers to access the data after the contest, creating a new library of algorithms that can be used in future studies and compared in an objective way.
“The only way cancer researchers can make informed decisions about the tools that they use is if the quality of algorithms is assessed objectively and independently,” said Amit Deshwar, a graduate student in Morris' group.
All participants who present a final algorithm will be co-authors on a Challenge overview paper, which will be submitted to Nature Biotechnology, a leading journal in the field and the official partner of the Challenge. Top performers will also receive travel awards and speaking invitations at the 2016 DREAM Conference, the 2016 Sage Congress or a similar event. Finally, the overall winning algorithms will be applied to real cancer data, including a subset of up to 2,500 whole genome tumour sequences.
“That’s a great rewards to think of: You can run your method on real patient data and ultimately make a contribution which could transform our understanding of cancer and how we treat it,” says Morris.
About Ontario Institute For Cancer Research
OICR is an innovative cancer research and development institute dedicated to prevention, early detection, diagnosis and treatment of cancer. The Institute is an independent, not-for-profit corporation, supported by the Government of Ontario. OICR research supports more than 1,600 investigators, clinician scientists, research staff and trainees located at its headquarters and in research institutes and academia across the Province of Ontario. OICR has key research efforts underway in small molecules, biologics, stem cells, imaging, genomics, informatics and bio-computing.
About Donnelly Centre for Cellular and Biomolecular Research
Donnelly Centre is an interdisciplinary research institute at the University of Toronto where scientists from diverse fields make discoveries into the fundamentals of biology and use these insights to improve human health. Founded in 2005 with the investment from the government of Canada and private sector contributions, the Centre has quickly established itself as a major international hub for biomedical research. Today the Donnelly Centre houses 35 principal investigators and 500 trainees and staff, who explore biology at all levels - from genes to proteins to cells. With partnerships that include the biotech sector and major Toronto hospitals, the Centre is well positioned to turn basic discovery into tangible advances in medicine.
About Sage Bionetworks
Sage Bionetworks is a nonprofit biomedical research organization, founded in 2009, with a vision to promote innovations in personalized medicine by enabling a community-based approach to scientific inquiries and discoveries. In pursuit of this Mission, Sage Bionetworks is working with others to assemble an information Commons for biomedicine that (1) is supported by an open compute space Synapse , (2) supports open research collaborations and innovative DREAM Challenges, and (3) empowers citizens and patients with the tools to partner with researchers and share their data through Sage’s BRIDGE platform in order to drive the research studies that matter most to them.
About DREAM Challenges
The Dialogue on Reverse Engineering Assessment and Methods (DREAM) Challenges pose fundamental questions about systems biology and translational medicine. A. Califano (Columbia University) and Gustavo Stolovitzky (IBM Research and the Icahn School of Medicine at Mount Sinai) founded the group in 2006. The DREAM Challenges, designed and run by a community of researchers from a variety of organizations, invite participants to propose solutions while fostering collaboration and building communities in the process. Expertise and institutional support are provided by Sage Bionetworks, along with the infrastructure to host challenges via their Synapse platform.
